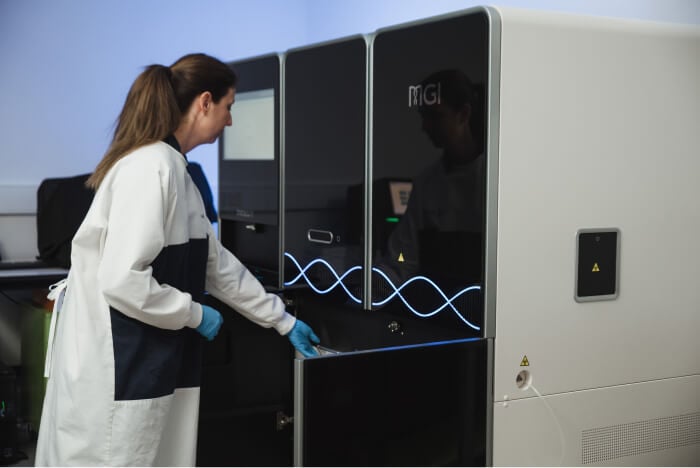
Heaps Good
Welcome to the South Australian Genomics Centre
Diverse Expertise
Agriculture, Biomedical, Environmental, MicrobialAny Scale
Range of next-gen sequencers to find a fit for projects of any sizeTailored
Customized solutions and leading-edge technologiesFast Turnaround
Dedicated team of genomic scientists and bioinformaticiansGet in touch
Located in the iconic SAHMRI building in the heart of Adelaide's Biomed City and at Flinders University (FHMRI).

.png?width=300&name=Copy%20of%20SA%20Omics%20Framework_Social%20Post%20(3).png)
.png?width=300&name=Untitled%20design%20(3).png)

Xenium in situ Analyser
Visualise 100s of RNAs within cells and tissues with off the shelf and custom probe sets for fresh frozen and FFPE tissues.
.png?width=300&name=Copy%20of%20Parse%20V3%20Kit_For%20Website%20(1).png)
Parse Evercode V3 NOW AVAILABLE
The new kit offers improved sensitivity, a faster workflow and more fixation capabilities.

Chromium GEM-X is HERE
Updates to the reagent chemistry and microfluidics for both the 3’ Single Cell Gene Expression and the 5’ Immune Profiling solutions.
Support from Sample through to Analysis
Genomics, Library Preparation, Single Cell, Spatial Transcriptomics, Bioinformatics Pipelines, Data Visualizations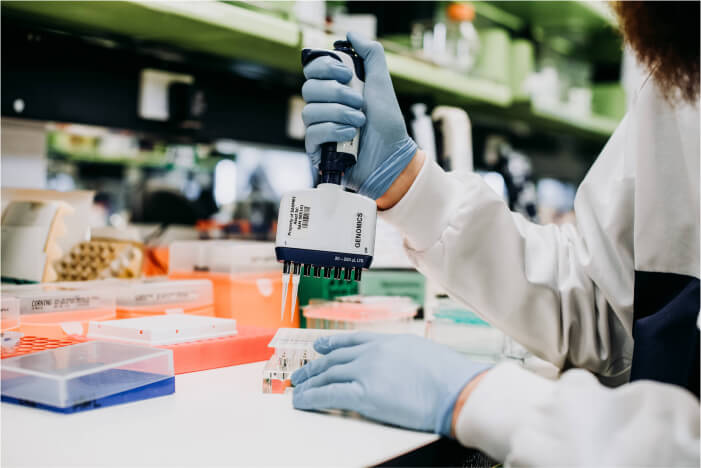
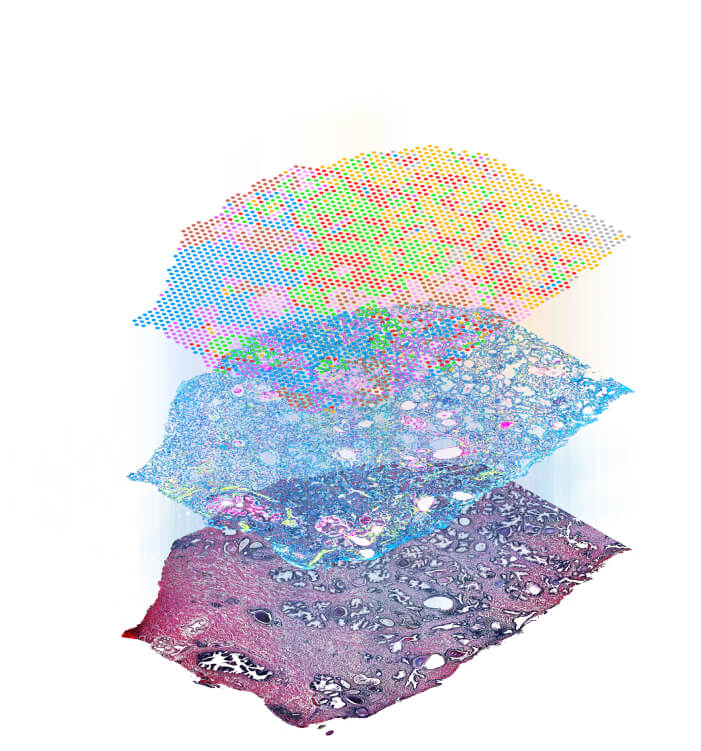

Xenium (10X Genomics)
Stereo-Seq (BGI STOmics)View PLATFORMS


Proudly Supported by

Support for Australian Biotech
We leverage our Genomics infrastructure to offer to end-to-end support for local industry partnersWorld-Class Research Capabilities in SA
South Australia is home to a range of state-of-the-art infrastructure to support basic through to translational research.
The SAGC can offer support with grant applications, project collaborations and connections with genomics industry.
Feedback from Clients

”The SAGC spatial transcriptomics service is a one stop shop. Wetlab experts made our experiment happen, and the computational service got us to a point that data analysis was possible without any complex coding. Landmark first spatial study of CTE would not have been possible without SAGC support.”

Acknowledgment of Country

The SAGC is located on Kaurna Country. We pay respects to the Kaurna people of the Adelaide Plains and to all Aboriginal and Torres Strait Islander people. We are committed to embracing knowledge and culture as we continue our working journey to incorporate Aboriginal health research across all of our themes and further reconciliation. The SAGC staff, Scientific Advisory Committee and Steering Committee are committed to ensuring that Indigenous Genomics is conducted in a culturally appropriate manner with Indigenous governance oversight.
"Tirki nguta tappa tangka marninithi"
(Learning and knowledge are the pathway to alter the mind)
- Uncle Lewis Yerloburka O'Brien, Kaurna Elder











